Regulation of Gene Expression in Eukaryotes
MCAT Biology - Chapter 2 - Section 3.1 - Gene Expression - Regulation of Gene Expression in Eukaryotes
- Home
- »
- MCAT Masterclass
- »
- Biological and Biochemical Foundations of Living Systems
- »
- Biology
- »
- Regulation of Gene Expression in Eukaryotes – MCAT Biology
Sample MCAT Question - Regulation of Gene Expression in Eukaryotes
How are eukaryotes able to produce multiple protein products from one gene?
a) Gene amplification
b) Binding of transcription factors
c) Histone methylation
d) Alternative splicing
D is correct.
Alternative splicing is a process by which different combinations of nucleotide sequences can be kept or removed in a pre-mRNA molecule. This process will increase diversity in protein products because it allows eukaryotes to produce multiple protein products from one gene.
Gene Expression: Prokaryotes vs Eukaryotes
Gene expression is the process of using the information found in genes to create gene products, i.e. proteins. This process is much more complicated in eukaryotes than in prokaryotes because eukaryotic mRNA transcripts are mostly monocistronic, meaning that they code for only one single protein product. Prokaryotes, by contrast, produce polycistronic mRNA transcripts, each of which code for multiple protein products. Thus, if a prokaryote needs to carry out a particular biochemical pathway involving multiple proteins, it might only need to produce one mRNA transcript. Eukaryotes, on the other hand, will need to produce separate mRNA molecules, one for each protein. This latter approach allows eukaryotes to express different combinations of proteins more efficiently, a fact which is especially important for multicellular organisms with specialized tissues and cell types. However, it also means that transcription needs to be more rigorously regulated. As a result, eukaryotes have evolved several different ways to regulate transcription and gene expression.
This MCAT post will discuss each of these regulatory mechanisms and how they allow the organism to transcribe and express the right gene at the right time.
Histone Acetylation
One method of transcriptional regulation in eukaryotes is the manipulation of chromatin structure. Recall that chromatin is DNA wrapped around histone proteins. One way of altering chromatin structure is histone acetylation, the process by which acetyl groups are added to, or removed from, on histone proteins. This process can alter the accessibility of DNA by RNA polymerase, which will alter the ability of the cell to transcribe genes. Interestingly, histone acetylation is involved in several types of lung diseases such as asthma, by regulation of inflammatory genes. Moreover, research has shown that histone acetylation can contribute to cancer development as well.
Histone Methylation
Another method of transcription regulation is by altering histone structure through histone methylation. In histone methylation, instead of adding or removing acetyl groups from histone proteins, methyl groups are being added or removed. This process can result in either an increase or a decrease in gene expression. It is important to note that histone methylation is different from DNA methylation. DNA methylation typically silences gene expression by adding methyl groups to DNA.
Transcription Factors
A third way of altering gene transcription levels is through the use of transcription factors. Transcription factors are proteins that can bind to DNA sequences called promoters or enhancer regions. When transcription factors bind to DNA, they can either decrease or increase gene expression. Figure 1 shows an example of transcription factors. Upon binding the promoter region, the transcription factors will increase the affinity of RNA polymerase for the promoter. By doing so, gene transcription will be increased.

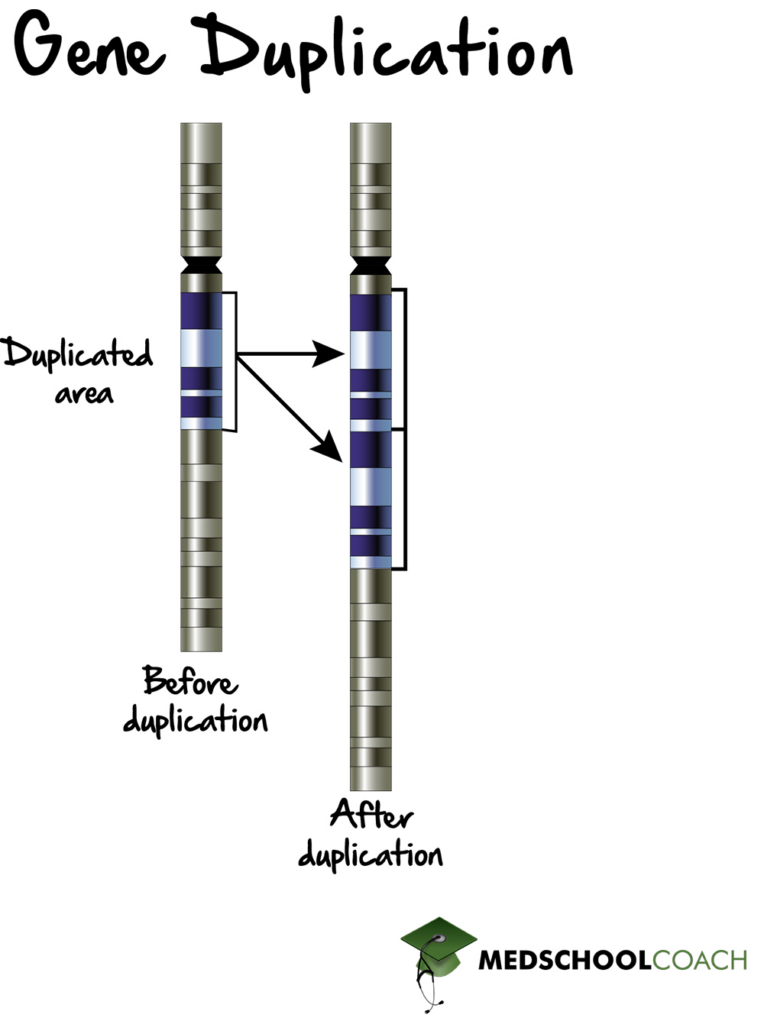
Gene Duplication
Gene duplication, also known as gene amplification, involves duplicating a region of DNA (Figure 2). If the duplicated DNA region includes both a promoter and as well as the entire coding region, this can lead to increased protein expression because now there are twice as many mRNA molecules and proteins being produced. It is important to note that gene duplication does not occur very often, and may even be a result of errors in DNA replication. Also, if the duplicated genes are oncogenes, which are cancer-causing genes, gene duplication can cause cancer. Some such cancers that can be caused through the amplification of oncogenes are breast cancer, cervical cancer, gastric cancer, or even lung cancer.
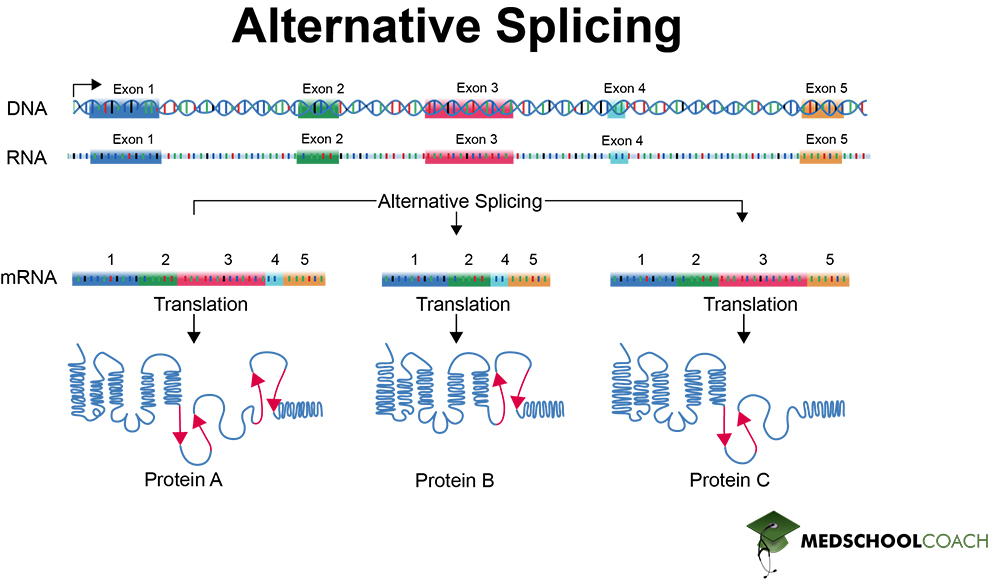
Alternative Splicing
Alternative splicing refers to the fact that different combinations of introns and exons can be capped or removed in a pre-mRNA molecule. Figure 3 illustrates how alternative splicing works. One DNA sequence is transcribed to pre-mRNA and then spliced in several different ways to produce many different protein products from the same gene. By having the ability to produce multiple protein products from one gene, alternative splicing increases the efficiency of gene expression in eukaryotes.
Non-Coding RNAs
Non-coding RNAs are RNA molecules that do not code for proteins. There are several types of non-coding RNAs that are involved in transcriptional regulation. MicroRNA, for instance, are short RNA fragments that can hybridize to complementary sequences on mRNA, thus blocking the translation of that gene and silencing gene expression. Another type of non-coding RNA involved in transcriptional regulation is small nuclear RNA, or snRNA. snRNA, along with several proteins, are the constituent components of spliceosomes, which are the complexes that catalyze mRNA splicing.
Explore More MCAT Masterclass Chapters
Take a closer look at our entire MCAT Masterclass or explore our Biochemistry lessons below.

One-on-One Tutoring
Are you ready to take your MCAT performance to a whole new level? Work with our 99th-percentile MCAT tutors to boost your score by 12 points or more!
See if MCAT Tutoring can help me
Talk to our enrollment team about MCAT Tutoring

MCAT Go Audio Course
Engaging audio learning to take your MCAT learning on the go, any time, any where. You'll be on the way to a higher MCAT score no matter where you are. Listen to over 200+ lessons.
MCAT Practice Exams
Practice makes perfect! Our mock exams coupled with thorough explanations and in-depth analytics help students understand exactly where they stand.
MCAT Prep App
Access hundreds of MCAT videos to help you study and raise your exam score. Augment your learning with expert-created flashcards and a question banks.